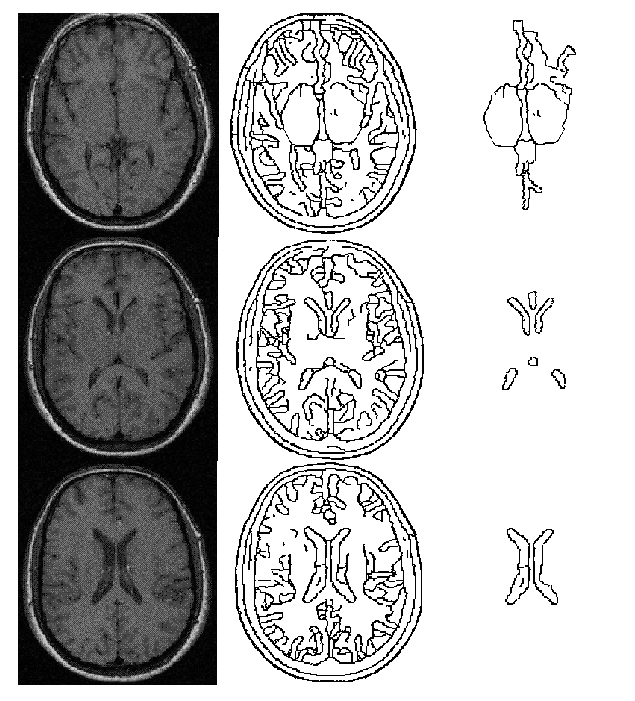
The object of this example was to use the techniques described in this paper to provide a labelling of the structures near the midline of a three slice MR data set. The input MR slices are shown in the left hand column of fig. 3. These were initially segmented using a standard Canny operator with hysteresis. For each slice the resultant edge points were then passed into the shape extraction process. The resultant boundary sections produced by this process are shown in the middle column of fig. 3. This process produced a large number of individual objects. To simplify the labelling process only a small number of these were considered as potential anatomical structures; in each slice only objects with an area above a certain size and with a centre of gravity within a specified distance from the middle of the image were considered as potential anatomical structures. Information about these structures was passed to the model environment for labelling. The boundaries for these objects are shown in the right hand column of fig. 3. Note that both skeleton and boundary representations exist for the objects shown in figure 3.
Information about the area, position, and direction of the skeleton for each object was passed to the model environment. Each object passed caused an individual frame to be generated to represent that object. An attempt was then made to match each image object to every midline structure stored within the model by comparing information about the size, position, and skeleton direction of each image object with the model object. This process returned a similarity value between 0 and 1. Any match with a value greater than a user defined threshold of 0.85 was considered a "good match". The value 0.85 was an arbitrary value. The choice of threshold is important; too high and there is a possibility of rejecting a correct match; too low and there is a possibility of a large number of incorrect matches been processed. Each good match caused a new viewpoint to be generated with the frame representing the image object used as the assertion for this viewpoint. An INSTANCE-OF link was generated connecting the frame representing the image object to the anatomical structure it was matched to. This processes allowed individual image objects to be classed as instances of multiple anatomical structures, each classification forming the basis of a separate viewpoint.
Merges between any two individual viewpoints were poisoned if the spatial relationships of the labelled image objects conflicted with the spatial relationships held within the model. Since individual image objects are connected to the inheritance network (via the INSTANCE-OF links), information regarding the expected spatial relationships of two image objects can be inferred from the spatial adjacency graph and the part-hierarchy.
Viewpoints were also poisoned if they had the same image object labelled as two different anatomical structures. Although it was not used in this example, each viewpoint contained a confidence measure for each labelling of an image object with an anatomical structure present in that viewpoint. These confidences are the result of the individual matching processes.
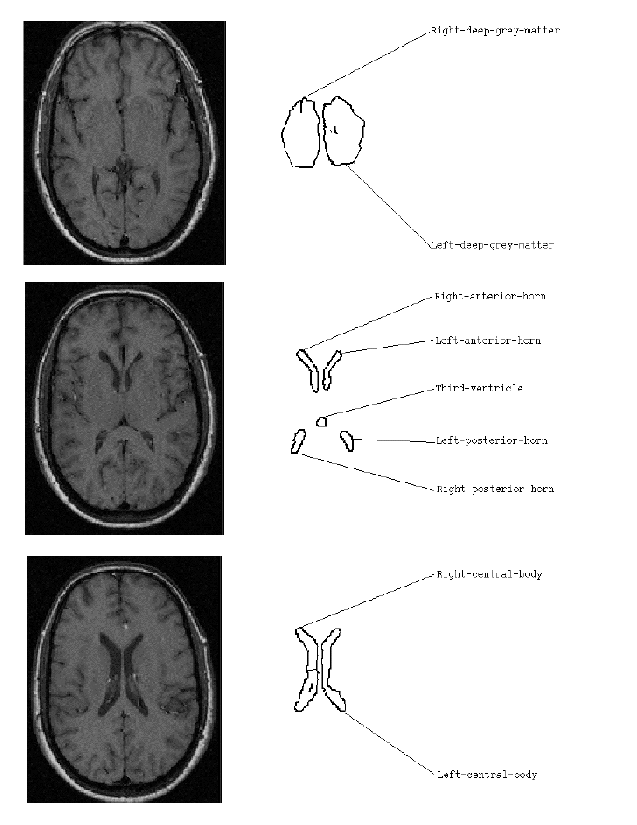
Each viewpoint only had image-to-model matches which did not conflict with the spatial relations held within the model. A series of rules then investigated the properties of each viewpoint to see how many image-to-model matches were present. Fig. 4 shows the image-to-model matches for the viewpoint with the most matches. The resultant image to model matches were all correct with one exception. The object labelled as "third-ventricle" is actually the "pineal recess". The incorrect match occurred since the image object's shape was similar to that of the shape of the third ventricle stored in the model, and its spatial relationships to the other labelled features did not conflict with those specified in the model for the third-ventricle. It is important to note that every viewpoint has an "allowable interpretation". The criteria used here to decide upon the viewpoint with the "best" interpretation was purely the number of image-to-model matches. There is no guarantee that this will always be the best interpretation. However, if the image objects produced correspond well to anatomical features then the number of matches is a good first estimate at an interpretation.